Project brief
The identification of progression biomarkers with Alzheimer’s disease is a fundamental component for identifying the molecular bases of the disease. A previous study has identified a genetic variant of the “number of copies” type in the SIRPB1 gene in 30% of the general Spanish population. Murine models have linked the action of the SIRPB1 protein with the ability to eliminate beta-amyloid deposits. The aim of our study is to determine to what extent the presence in an Alzheimer’s patient of this variant in the SIRPB1 region could be affecting the phagocytic capacity of the microglia and alter the progression of cognitive impairment.
This is a retrospective cross-sectional genetic analysis study in which no intervention will be performed on the patients. Patients with a possible or probable diagnosis of Alzheimer’s who have > 3 months of evolution records from the moment of diagnosis will be evaluated.
Fundamentals
In 2014, our group identified a genetic variant that affected the expression of the SIRPB1 gene. This is an insertion/deletion variant in the coding sequence of the gene, which has a frequency of 30% in the general Spanish population. Alzheimer’s disease that have > 3 months of evolution logs from the time of diagnosis.
It has been documented that SIRPB1 regulates neutrophil migration by playing a unique role in regulating the inflammatory response. Likewise, SIRPB1 has been shown to act as a phagocytic receptor in microglia in the model of transgenic mice for Alzheimer’s disease (APP mice) and in the model of experimental autoimmune encephalomyelitis (EAE mice). In these experimental models, the activation of SIRPB1 in microglia cells increased the uptake of microspheres, neuronal remains and beta-amyloid protein.
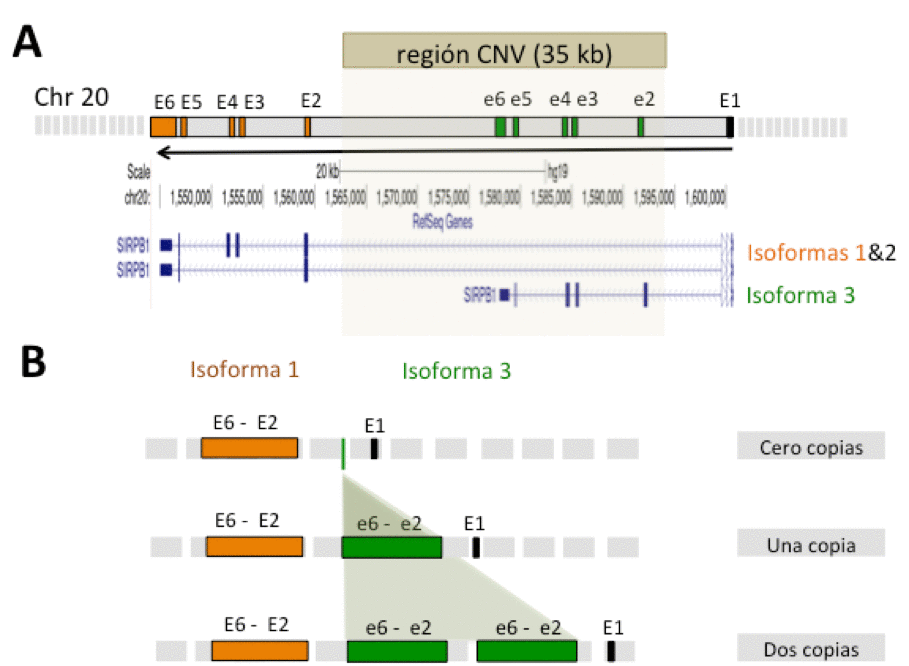
Figure 1.- Structure of the genomic region of the SIRPB1 locus. In A, the structure of the three isoforms (1, 3 and 3) of the SIRPB1 gene and the delimited region pro in CNV-SIRPB1 are shown. In B, the different alleles of CNV-SIRPB1 that determine the existence of zero, one or two copies of the CNV region are shown schematically.
To date, it has not been considered to establish a relationship between the presence of functional variants of SIRPB1 and their potential implication in the age of onset and progression of the disease. To substantiate this, it is necessary to obtain CNV-SIRPB1 genotype information in genomic DNA samples by monitoring progressive data from individuals with cognitive impairment. This will unequivocally allow the establishment of whether or not there is a correlation between the CNV copy number and its role in AD. The presence of CNV in the region of SIRPB1 would alter the phagocytic capacity of the microglia and this would modify the age of onset of the disease and alter the progression of cognitive impairment.
Starting hypothesis
The presence of CNV in the region of SIRPB1 alters the phagocytic capacity of the microglia and this fact modifies the age of onset of the disease and alters the progression of cognitive impairment.
Objectives
Generate a collection of 250 patient samples and digitize the epidemiological parameters related to the study.
Extract the genetic material from the collection samples and determine the variation in the number of copies of the SIRPB1 gene (CNV-SIRPB1).
Carry out the CNV-SIRPB1 association study with the clinical variables collected in the database: Fundamentally, the initial ages and degree of clinical manifestation of the pathology.
Disseminate research results.
Collection of biological material
Alzheimer’s patients with at least 3 months follow-up from the date of diagnosis who altruistically wish to participate in the study will be provided with an informed consent form. Only epidemiological and clinical evolution data relevant to this study will be taken. The data will be linked exclusively by means of an alphanumeric code on a sticker.
A sample of 1-3 ml of peripheral blood will be taken during a routine control in vials with EDTA that will be frozen at -20ºC.
DNA will be obtained from each of the samples to be analysed using specific commercial kits (QIAamp DNA Blood Mini Kit). The concentration will be determined by PicoGreen (Quant-iT™ PicoGreen® dsDNA Assay Kit) and its integrity by agarose gel electrophoresis.
Anonymized DNA samples will be conveniently stored in the collection with registration number C0004388 of the National Registry of Biobanks (Instituto de Salud Carlos III), for which Dr. Royo is responsible. This collection is located in the facilities of the Department of Biochemistry of the Faculty of Medicine in accordance with the specifications described in Royal Decree 1716/2011.
GERALD Consortium
José María García-Alberca 1
Silvia Mendoza 1
Esther Gris 1
Paz De la Guía 1
María López de la Rica 1
José Luis Royo 2
Laura González 7
José Manuel Cruz-Gamero 7
Emilio Alarcón-Martín 7
Luis Miguel Real 7
Irene Gonzalez 7
Maximiliano Ruiz 7
Armando Reyes-Engel 7
María José Bravo 7
Lidia Lopez-Gutierrez 7
María Dolores Nieto 3
Concepción Urbano 3
Rocío Jiménez-Sánchez 3
Natalia García-Casares 4
Macarena Luque 5
María García-Peralta 5
Rosario Carrillejo 6
María del Carmen Furniet 7
Lourdes Rueda 7
Ana Sánchez-Fernández 8
Tomás Mancilla 8
Isabel Peña 8
Nuria Pareja 9
Olga Ocejo 10
Javier Torrecilla 10
Carmen Zafra 10
1. Research Center and Memory clinic. Instituto Andaluz de Neurociencia (IANEC),
Málaga, Spain
2. Departamento de Especialidades Quirúrgicas, Bioquímica e Inmunología. Facultad de
Medicina. Universidad de Málaga. Málaga, Spain
3. Hospital Hermanas Hospitalarias del Sagrado Corazón, Málaga, Spain
4. Departamento de Medicina, Facultad de Medicina, Universidad de Málaga, Málaga,
Spain
5. Centro Residencial Élite, Málaga, Spain
6. Asociación Criptana de Enfermos de Alzheimer, Campo de Criptana, Spain
7. Asociación de Familiares de Alzheimer de Archidona, Archidona, Spain
8. Residencia DomusVi Fuentesol, Alhaurín de la Torre, Spain
9. Asociación de Familiares de Enfermos de Alzheimer de la Axarquía, Vélez-Málaga,
Spain
10. Centro Residencial Almudena, Rincón de la Victoria, Spain
Lead investigators:
Dr. José María García-Alberca (IANEC)
Dr. José Luis Royo (UMA)
